Transitioning from Bulk RNA-Seq to Single-Cell RNA Analysis (scRNA-seq)
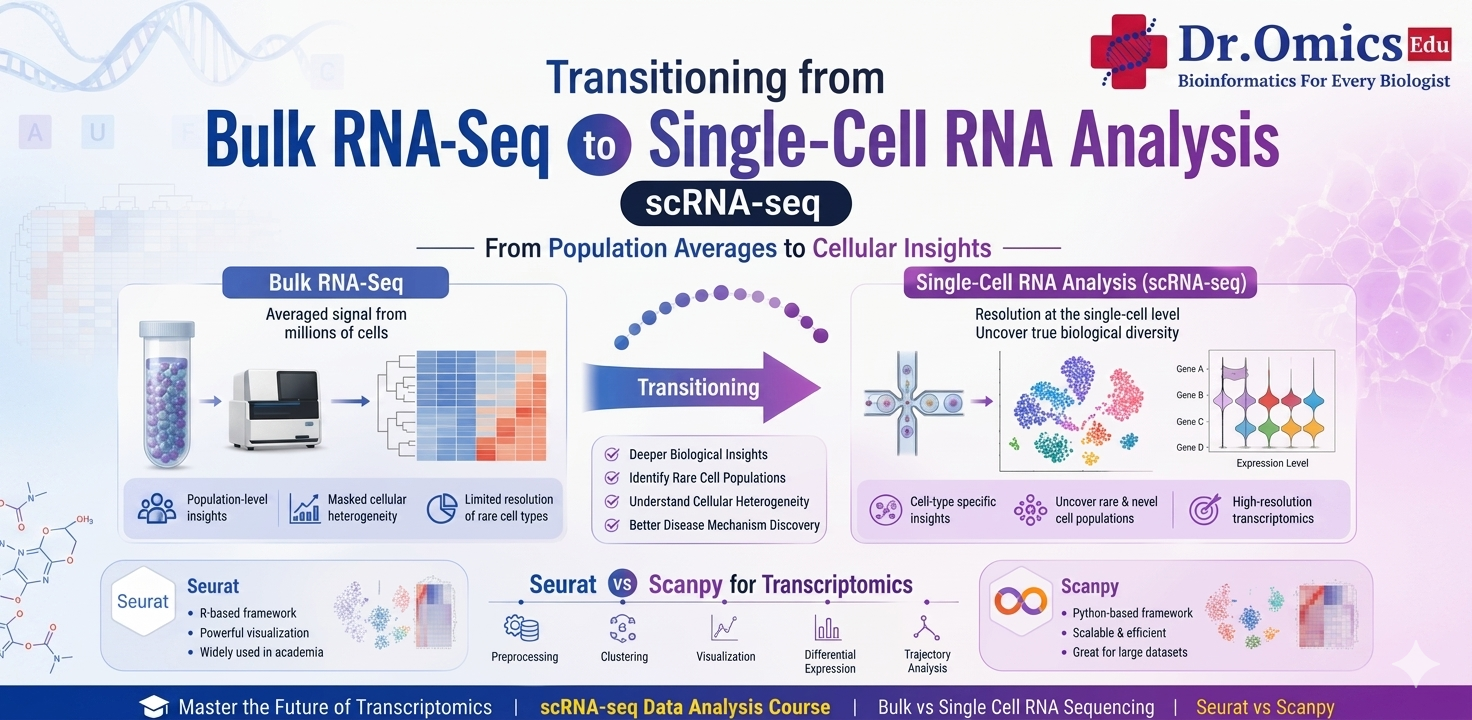
RNA sequencing has become an essential technique for studying gene expression. Traditional bulk RNA sequencing measures the average gene expression across thousands or millions of cells in a sample. While it is useful for identifying overall gene expression changes, it cannot reveal differences between individual cell types. This limitation has led researchers to adopt single-cell RNA sequencing (scRNA-seq), which allows gene expression analysis at the level of individual cells.
Understanding the difference between bulk vs single cell RNA sequencing is important when transitioning to modern transcriptomics approaches. Bulk RNA-seq provides an average signal from all cells, which can hide important biological variation. In contrast, scRNA-seq captures gene expression in each cell separately, making it possible to identify different cell populations, rare cell types, and cellular states within a tissue.
The workflow of scRNA-seq data analysis includes several key steps such as quality control, normalization, dimensionality reduction, clustering of cells, identification of marker genes, and cell type annotation. These steps help researchers understand how different cells behave and interact within complex biological systems.
Two of the most commonly used tools for single-cell transcriptomics are Seurat and Scanpy. Seurat is an R-based toolkit widely used for scRNA-seq analysis and provides a comprehensive framework for clustering, visualization, and integration of datasets. Scanpy is a Python-based tool designed for scalable analysis of large single-cell datasets and integrates well with machine learning workflows. When comparing Seurat vs Scanpy for transcriptomics, the choice often depends on whether the researcher prefers working in R or Python.
As single-cell technologies become more common in genomics research, many scientists are now looking for structured training such as an scRNA-seq data analysis course. These courses help researchers learn the fundamentals of single-cell transcriptomics, data preprocessing, clustering methods, and biological interpretation.
Overall, transitioning from bulk RNA-seq to single-cell RNA analysis allows researchers to explore cellular heterogeneity and gain deeper insights into biological systems. With tools like Seurat and Scanpy and proper training in scRNA-seq workflows, scientists can unlock the full potential of single-cell transcriptomics.